TPJ article: Genomics Reveal a Complicated Past and Suggest a Possible Future for Nicotiana benthamiana
Genomics Reveal a Complicated Past and Suggest a Possible Future for Nicotiana benthamiana
Luiz A. Cauz-Santos, Steven Dodsworth, Rosabelle Samuel ,Maarten J. M. Christenhusz, Denise Patel, Taiwo Shittu, Aljaž Jakob, Ovidiu Paun, Mark W. Chase
Genomic insights into recent species divergence in Nicotiana benthamiana and natural variation in Rdr1 gene controlling viral susceptibility
Nicotiana benthamiana is one of the most widely used model organisms in plant biology. But the story of this little plant’s rise to prominence is actually quite serendipitous. Benjamin Bynoe, a surgeon aboard the HMS Beagle who worked with Darwin in the Galapagos, was the first person to collect this species from the north-west coast of Australia in 1839. However, the plants currently grown in laboratories are descended from a single collection of seeds made in the late 1930s by Professor John Cleland from the University of Adelaide, at a site in the Tanami Desert. In 1939, Cleland sent some of these seeds to Professor Thomas Goodspeed at the University of California, Berkeley. Goodspeed propagated them and sent seed to other scientists, who quickly realized N. benthamiana was all but incapable of combating diseases.
Because of this incredible susceptibility to pathogens, especially viruses, N. benthamiana has become a popular model organism for plant pathology research. It wasn’t until 2004, however, that researchers discovered the genetic basis of this seemingly maladaptive trait: a knockout mutation in RDR1, encoding an RNA-dependent RNA polymerase (Yang et al., 2004) that functions in RNA silencing, an integral aspect of immune responses against RNA viruses (Xie et al., 2001). But what possible evolutionary advantage could there be for a severely weakened immune system?
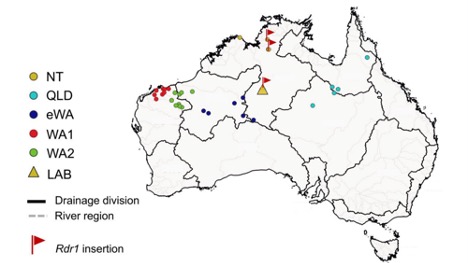
A team addressing this question in 2015 discovered that only one of their N. benthamiana accessions contained the insertion that knocks out RDR1. They also noticed that this mutation was associated with larger seeds, faster germination, and more vigorous seedlings. So, they hypothesized that strains carrying a mutated Rdr1 allele, including the laboratory isolate (LAB), are desert extremophiles that have essentially abandoned viral resistance in favor of precocious growth. However, an article published in issue 111:1 of The Plant Journal offers another look at the ecology of this evolutionary peculiarity and comes to a different conclusion. Cauz-Santos and colleagues examined the relatedness of 36 different N. benthamiana accessions from across Australia. They concluded that LAB is more closely related to accessions found far north of Cleland’s original collection site, which grow in areas much wetter than the Tanami Desert. They also provided evidence that N. benthamiana should be split into five separate species. Of these proposed species, the Northern Territory (NT) clade, which contains LAB, is the most divergent, and the Rdr1 mutation is found only within this population (Figure 1).
The authors go on to show that although LAB is the fastest to germinate, it is not the fastest to flower and set seed. Since it is also most closely related to other accessions from more moderate environments, the authors refute the previous claims that LAB is an extremophile optimized for rapid generation cycling. It remains unclear why members in the NT clade would have jettisoned viral defense, although the authors suggest that if there is little opportunity for viral exchange within the species, a loss of virus resistance might not be detrimental. Other researchers have suggested that certain traits, potentially including the Rdr1 knockout, were selected by First Nations peoples in Australia, who were using it as chewing tobacco, raising important ethical points about indigenous intellectual property rights. Although some mystery still surrounds the traits that make N. benthamiana such an attractive model plant for research, these findings are a significant advance in understanding its evolutionary history and in establishing a more accurate taxonomic consensus.





