TPJ Editor choice: Arabidopsis callus-grafts: A system to study symplasmic intercellular macromolecular transport and gene silencing spread
Arabidopsis callus-grafts: A system to study symplasmic intercellular macromolecular transport and gene silencing spread (https://doi.org/10.1111/tpj.16326)
Local call or long-distance call: studying cell-to-cell transport by Arabidopsis callus grafting
Connections between plant cells were discovered more than a century ago by the Austrian botanist Eduard Tangl. He proposed that they serve for communication and transport between cells. Today it is clear that these connections - termed plasmodesmata - facilitate the movement of molecules such as mRNA and proteins between cells. Recently, it was demonstrated that it is possible to create stable gene-edited, transgene-free plants by grafting a wild type shoot stock to a rootstock transformed with a CRISPR/Cas9 construct. Fusions of Cas9 and guide RNA transcripts to a tRNA-like sequence motifs move the RNA from root to shoot and create inheritable mutations in the shoot.
How is this RNA transport achieved between cells over short distances? Studying symplasmic transport of macromolecules between cells in plants independent of transport in the vasculature is difficult. Hasbioğlu et al. describe a callus-grafting system to study local macromolecule transport without the presence of a vascular system. For this, they co-cultivated calli to form a chimeric tissue. They used YFP as a mobile protein expressed in the donor callus and Cherry fused to histone H2B as immobile protein expressed in the recipient tissue. At the attachment sites of the calli, they detected cells with both fluorescent proteins, suggesting that YFP was transported between the cells. Calli from two different ecotypes, as well as calli derived from different plant organs, could form symplasmic connections. Cryo-sections of the connection sites showed that the plasmodesmata were labelled with the marker for complex plasmodesmata from one side and with a general marker for plasmodesmata from the other side, suggesting that plasmodesmata are formed at the chimeric junction site. The authors then tested the mobility of macromolecules other than YFP through these plasmodesmata, confirming the mobility of the transcription factor, KNOTTED 1 (KN1), which is known to move from cell-to-cell by regulated transport.
The unorganised nature of the calli and their loose structure limited the number of observed fluorescent protein mobility at the graft junction, making the authors speculate the events were under-counted. Therefore, they went on to use small interference RNA (siRNA) silencing spread, which depends on symplasmic transport and is amplifying, so that only one connection event at the calli junction site should be enough to silence gene expression in the whole callus. The authors used two gRNA-TLS (tRNA-like Sequence) expressed in the recipient callus to target VENUS, and a control gRNA targeting another locus. In the negative control, VENUS signal was still visible in the recipient callus expressing 35S:H2B-VENUS but abolished in the experimental setup containing VENUS-targeting siRNA, suggesting that the silencing spread was established throughout the recipient callus.
As well as proteins, mRNAs are known to be transported across cells and tissues via the symplasm. The mRNA of PEROXIN 11D (PEX11D) is predicted to be mobile, but the protein localises to the ER and peroxisomal membranes and is therefore unlikely to move itself. Nevertheless, when Hasbioğlu et al. expressed YFP-PEX11D in the donor callus, the authors were able to detect the protein reporter in the recipient calli at the grafting sites, suggesting that the mRNA moved between the calli and that the method was feasible to detect this movement. According to Frank Machin, corresponding author of the study, the callus grafting method can answer the most pressing question in the field of mobile mRNA research, “which mRNAs can be detected moving across callus grafts?” It will give insights into the properties of specific mRNAs and help to identify sequence motifs that can indicate if a particular mRNA is cell-to-cell mobile.
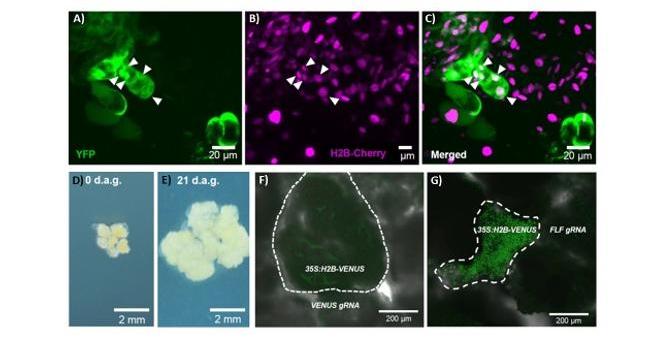
Figure 1: Callus grafting for analysing symplasmic transport.
A-C) YFP is mobile at the grafting sites between a callus expressing YFP and a callus expressing H2B-Cherry; arrow heads mark the nuclei that show YFP and Cherry signal; YFP channel in green (A), Cherry channel in magenta (B), merge (C).
D-E) Assembly of calli on callus induction medium; the recipient callus is placed in the center (0 days after germination (d.a.g), D) and the calli are grown until the cells attach at the junction site (E).
F-G) Graft between a callus expressing 35S:H2B-VENUS and a donor callus expressing gRNA targeting VENUS, which induces silencing of VENUS in the recipient callus (dashed line, F); no silencing is observed when FLOWERING LOCUS F is target by gRNA from the donor callus; modified from (Hasbioğlu et al., 2023).





