TPJ article: Once, twice, three times a genome – how the three sub-genomes of a water lily control leaf development
Wu, P., Zhang, L., Zhang, K., Yin, Y., Liu, A., Zhu, Y., Fu, Y., Sun, F., Zhao, S., Feng, K., Xu, X., Chen, X., Cheng, F. and Li, L.
The adaptive evolution of Euryale ferox to the aquatic environment through paleo-hexaploidization
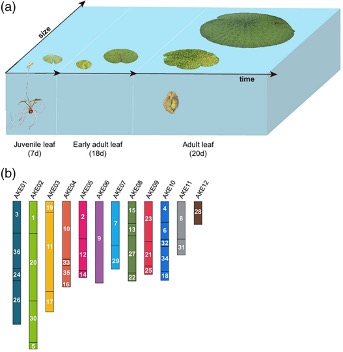
Figure 1. (a) Different developmental stages of E. ferox leaves. (b) The reconstructed chromosomes of the diploid ancestral genome of E. ferox. The numbers on the bars are the IDs of corresponding genomic blocks. AKE, ancestral karyotype of E. ferox. Figure adapted from Wu et al., 2022.
Once, twice, three times a genome – how the three sub-genomes of a water lily control leaf development
Nymphaea colorata and Euryale ferox are relatively closely related water lily species that look rather different: N. colorata develops relatively small leaves, whereas, E. ferox develops gigantic prickly floating leaves that can grow up to 2 m in diameter, much like the leaves of the Santa Cruz waterlily (Victoria cruziana). E. ferox leaves develop in a particular way: small peltate juvenile leaves grow on the water surface and transition into round adult leaves. When the adult leaf is about 18 days old, it disappears under water and wraps up like a ball. The leaf ball grows and then emerges from the water a few days later, when it unfolds and expands into a flattened leaf of maximum 100 cm within one day. For the following days, it continues growing (Figure 1a). Recently, the genomes of Nymphaea colorata and Euryale ferox were sequenced. Analysis of these genome sequences showed that E. ferox has undergone an ancient allo-hexaploidization event that N. colorata has not. In issue 110:3 of The Plant Journal, Peng Wu and colleagues investigated this genome triplication and tried to understand how it contributed to the development of leaves in E. ferox.
To get insight into the three sub-genomes of E. ferox, the researchers reconstructed the ancestral set of chromosomes of its diploid ancestor. To do this, they used the genome of N. colorata (14 chromosome pairs) as a reference to sort syntenic genomic fragments of E. ferox (29 chromosome pairs). Subsequently, they searched for break points: if two syntenic blocks were arranged consecutively on one genome, but not on the other, a break point was defined. If a break point was present in more than one copy of the syntenic fragment, it was considered to be inherited from the ancestral genome. Next, they aligned these genomic blocs along the genome of E. ferox and searched for associations between the blocks – basically the opposite of a break point: where the genomic blocks were arranged consecutively on the E. ferox genome for more than one copy, their association was considered inherited from the ancestral genome. The resulting associations fused the genomic blocks into 12 groups, suggesting that the ancestral genome of E. ferox before hexaploidization had 12 chromosome pairs (Figure 1b). With these 12 chromosomes as a reference, the authors sorted each of the copies of the syntenic fragments and reconstructed three versions of the 12 chromosome pairs, corresponding to the three sub-genomes that merged in the whole-genome triplication event.
The authors then investigated the phylogenetic relationships between V. cruziana, N. colorata, and the three sub-genomes of E. ferox by comparing syntenic orthologous genes. For each of the reconstructed ancestral chromosome copies, there was always one chromosome copy that was more closely related to V. cruziana, and each of these chromosomes belonged to the same sub-genome. However, when the authors evaluated the number of genes that were retained from each sub-genome, they did not find a difference.
They next measured gene expression of paralogous gene pairs to identify the dominantly expressed gene for each of the paralogous genes. The dominantly expressed genes were distributed evenly over the three sub-genomes. Hence, there appeared to be no genome-wide dominance of one of the sub-genomes in E. ferox that can explain the leaf phenotype.
Therefore, the authors performed RNA-seq on 11 different time points of developing leaves, as well as on the root, stem, flower, fruit, and seeds. To annotate organ- and time point-specific genes, the authors compared all the datasets to identify genes that are specifically expressed in the different organs or at specific time points during leaf development. When they evaluated the presence of these annotated genes on the different sub-genomes, they found that the sub-genome closest to V. cruziana was enriched in genes that were specifically expressed during the development of the adult underwater leaf (the leaf-ball stage). This suggests that this sub-genome of E. ferox contributed more to the adaptive evolution of the leaves, and could explain why E. ferox and V. cruziana have similarly large leaves.





